Big Data Department makes progress in supporting Microbiological Field Database and its Analysis System
The Big Data Department of CNIC and the Institute of Microbiology, CAS and other teams have made new progress in the construction of microbiology database and analysis system. The team has presented a method of mapping data from publicly available genomics and publication resources to the Resource Description Framework (RDF) and implemented a server to publish linked open data (LOD). These LOD with 62,168,127 semantic triplets. The resulted gcCov database demonstrates the capability of using data in the LOD framework to promote correlations between genotypes and phenotypes. These correlations will be helpful for future research on fundamental viral mechanisms and drug and vaccine designs. The research results have been published in the academic journal “mLife”.
In recent decades, coronaviruses (CoVs) have caused severe infectious diseases and posed a continuous global threat to public health. This has resulted in extensive research on novel human and animal CoVs and the rapidly increasing number of such publications. This rapid expansion in data has inevitably led to the great challenge of integrating diverse types of studies into a single searchable correlated source. Currently, available CoV databases mainly focused on genomic analysis (e.g., CovDB1 and ViPR2) or publications (e.g., LitCovid3), or mainly focused on SARS‐CoV‐2 (2019nCoVR4,5). However, these databases did not establish correlations between genomic data and other types of information (e.g., papers, patents, and antibodies).
With the ability to integrate distributed web resources into a knowledge base of shared ontologies and then analyze those data to identify underlying relationships between various entities, the semantic web is a promising solution for biomedical data integration. To analyze the relationship between massive data, this study designed a pipeline method to integrate data from different sources into the semantic web framework. Based on this method, the gcCov database was constructed and provides extensive information and relationships regarding CoVs using linked open data (LOD). The gcCov is the first and only CoV database published using LOD and based on a semantic web framework. It helps scientists to detect connections between the linked data and hence to discover new knowledge that would otherwise be hidden in the mass data. This type of database is an important tool for the increasing information needs of CoV research, given its ability to mine text and data from previous studies and to provide clues for current prevention and treatment strategies.
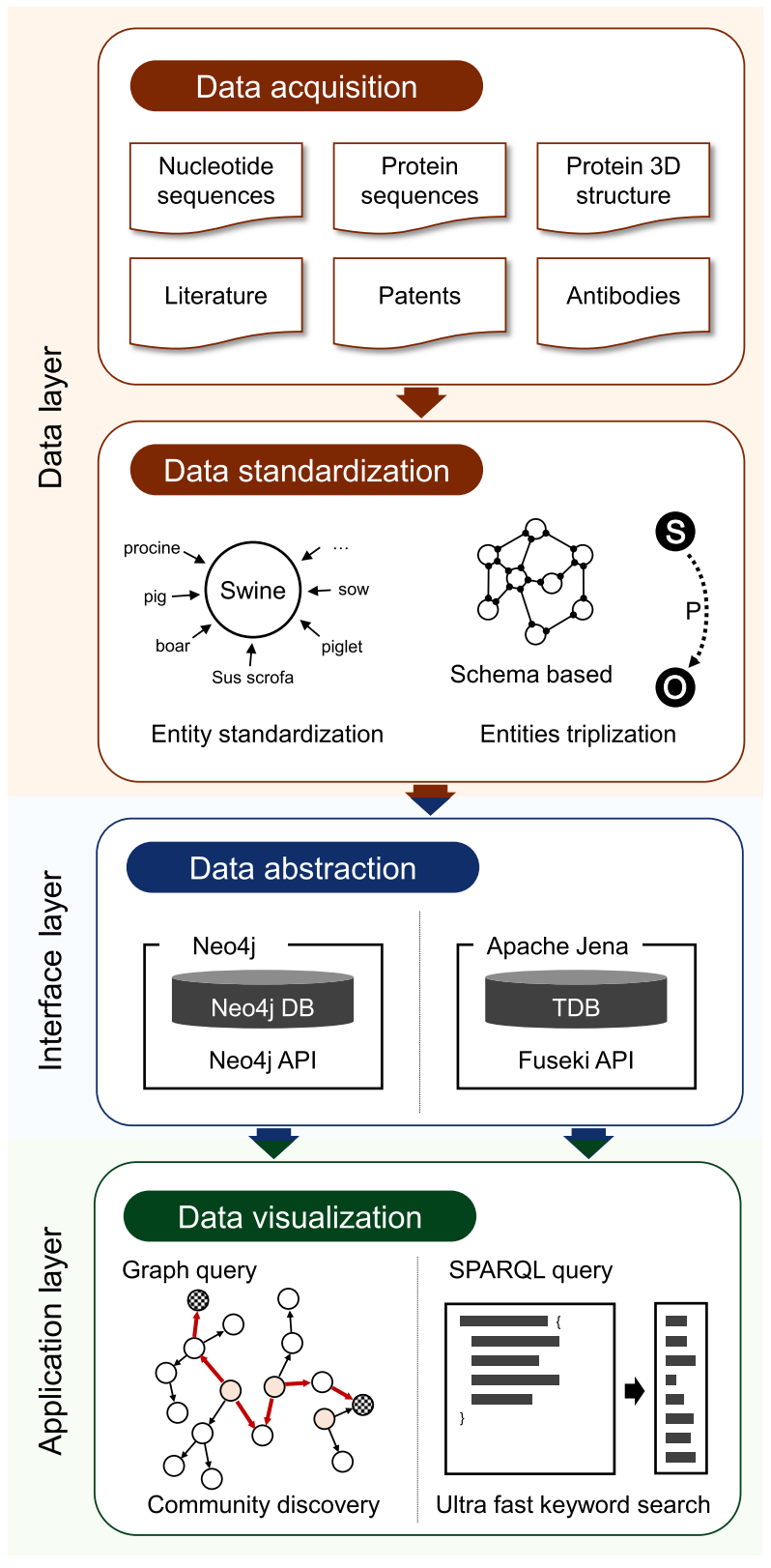
Figure 1. Pipeline of data processing and utilization.
For details, please contact Hu Chuan (huchuan@cnic.cn)
